Macrophages tolerization analyzed by high resolution plasmon imaging
Internship at UGA/CEA/CNRS lab Molecular Systems and nanoMaterials for Energy and Health (SyMMES)
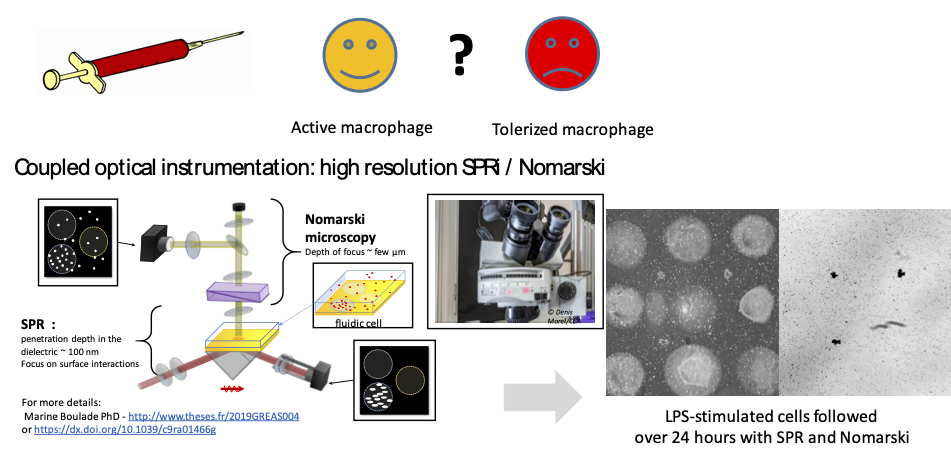
Scientific context and objective
Tolerization is a mechanism that appears after an episode of strong solicitation of the immune system. It results in the freezing of the immune system, which becomes less reactive to new infections. This mechanism of tolerization could be understood as a beneficial negative control of the inflammatory response, but its prolongation, as observed in some septic patients, can be deleterious and associated with an increased mortality.
The recognition of this state in clinical settings has not been properly addressed yet. For this reason, developing a methodology to allow an early identification of the tolerance state (before the occurrence of an infection) remains a relevant and challenging objective.
Instead of striving to detect molecular changes in the environment, as does by the classical approach, we want to develop an original paradigm by investigating the global phenotypical changes of the involved cells. For that we propose to evaluate the efficiency of a cell-based SPR approach to obtain a label-free readout of monocyte/macrophage tolerization state at cell level (i.e. identification, quantification and temporal study of tolerized cells).
Biosensor principle
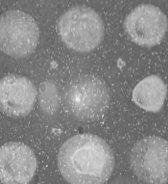
The proposed technology is based on cellular biochips studied with surface plasmon resonance imaging (SPRi), an optical imaging method widely used for the analysis of molecular interactions. This technique is particularly adapted for the measurement of variation in density of cells close to the chip surface. High-resolution SPR analysis (coupled with classical microscopy) should thus provide access to global morphological modifications (for example due to changes in adhesion properties), which may constitute the signature of the tolerization.
Trainee's work
The M1 trainee will receive training on biochip preparation and biosensors. She/he will then take charge of the SPR (HR) device previously developed in the laboratory, to study the changes over time of immune cells in chip culture.
This work involves cell manipulation, image processing, programming skills (instrumentation and/or image processing) knowledge in optics, biochemistry and surface functionalization. Obviously all these skills are not a prerequisite (the laboratory has them) but the interest for multidisciplinary work and its requirement is essential.
Thereafter (M2 internship), once the technique has demonstrated its ability to translate cell changes into measurable signals, the trainee will treat these signals, applying for example principal component analysis, to precisely define relevant signature of tolerization. Studies will begin in simple culture media and will gradually be performed on more and more realistic samples (blood).
This work will be done in collaboration with the Institute for Advanced Biosciences (IAB).
Background expected
master student in the Soft Nanosciences track, or student from Phelma Biomedical Engineering.
For more details
Marine Boulade PhD - http://www.theses.fr/2019GREAS004 and https://dx.doi.org/10.1039/c9ra01466g
Published on March 14, 2022
Updated on June 23, 2023